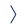
Interaction of hepatitis B viral proteins with host cell machineries
Hepatitis B virus (HBV) infection is a global public health problem, affecting more than 400 million people worldwide and placing them at high risk of developing chronic liver diseases including cirrhosis and hepatocellular carcinoma.
The key factor in viral persistence is covalently closed circular (ccc) DNA, an intracellular HBV replication intermediate that resides in the nucleus of infected cells as a non-integrated plasmid-like molecule and serves as a transcriptional template for HBV. Despite extensive research, the contribution of HBV proteins to the development and regulation of chronic infection remains poorly understood, and strategies for cccDNA degradation and elimination are limited.
Our main goal is to understand the role of the HBx protein in stimulation of HBV transcription and development of chronic hepatitis B. HBx is a multifunctional protein, and its biological functions are mainly dependent on pleiotropic protein–protein interactions. The role of HBx in redirection of the DNA-damage binding protein 1 (DDB1), the adaptor of Cullin-RING ubiquitin-E3 (CLR4E3) ligase to target cccDNA transcription restriction Smc5/6 complex-specific proteins for degradation was reported recently (Decorsière et al. 2016). Our group is working on structural characterization of potential complexes formed during degradation of this restriction factor.
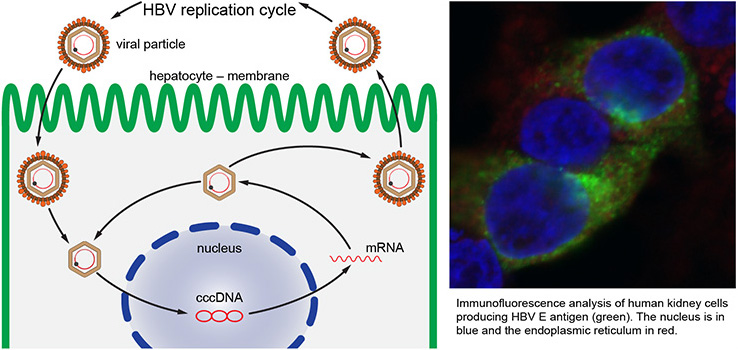
Another project focuses on processing of HBV precore protein precursor (HBe) during replication. HBe is processed stepwise in host cells, and its final product called p17 is released and serves as a clinical marker for disease severity. We analyze the factors influencing HBe processing and localization and study the role of host factors in HBe subcellular localization.
Project publications
- Zábranská H., Zábranský A., Lubyová B., Hodek J., Křenková A., Hubálek M., Weber J., Pichová I.: Biogenesis of hepatitis B virus e antigen is driven by translocon-associated protein complex and regulated by conserved cysteine residues within its signal peptide sequence. FEBS J. 2022, 289:2895-2914.
- Lubyová B., Tikalová E., Krulová K., Hodek J., Zábranský A., Hirsch I., Weber J.: ATM-Dependent Phosphorylation of Hepatitis B Core Protein in Response to Genotoxic Stress. Viruses. 2021, 13(12):2438.
- Langerová H., Lubyová B., Zábranský A., Hubálek M., Glendová K., Aillot L., Hodek J., Strunin D., Janovec V., Hirsch I., Weber J.: Hepatitis B Core Protein Is Post-Translationally Modified through K29-Linked Ubiquitination. Cells. 2020, 9(12):2547.
- Janovec V., Hodek J., Clarová K., Hofman T., Dostalík P., Froněk J., Chlupáč J., Chaperot L., Durand S., Baumert T. F., Pichová I., Lubyová B., Hirsch I., Weber J.: Toll-like receptor dual-acting agonists are potent inducers of PBMC-produced cytokines that inhibit hepatitis B virus production in primary human hepatocytes. Sci. Rep. 2020, 10:12767.
- Ramakrishnan D., Xing W., Beran R. K., Chemuru S., Rohrs H., Niedziela-Majka A., Marchand B., Mehra U., Zábranský A., Doležal M., Hubálek M., Pichová I., Gross M. L., Kwon H. J., Fletcher S. P.: Hepatitis B virus X protein function requires zinc binding. J. Virol. 2019, 93:e00250-19.
- Lubyová B., Hodek J., Zábranský A., Prouzová H., Hubálek M., Hirsch I.,
Weber J.: PRMT5: A novel regulator of Hepatitis B virus replication and an arginine methylase of HBV core. PLOS ONE. 2017, 12(10):e0186982.
People
Aleš Zábranský, Helena Zábranská, Michal Doležal, Vicent Llopis-Torregrosa, Karolína Štaflová, Kamila Clarová, Olena Berehovska, Romana Hadravová, Romana Cubínková

