Functional analysis of enzymes regulating Mycobacterium tuberculosis metabolism
According to the WHO estimates, one-third of the world’s population is latently infected with Mycobacterium tuberculosis (Mtb). However, the biology of Mtb that persists in infected people is still poorly understood. Mycobacteria utilize a whole range of stress response mechanisms and reprogram its metabolism during latent infection in response to the host environment. This metabolic adaptation is a very complex and dynamic process leading to the formation of the highly persistent dormant state of Mtb.
We perform structural and functional characterization of enzymes involved in central carbon metabolism, biosynthesis of nucleic bases and stress signaling; in particular, we focus on phosphofructokinases A and B (EC 2.7.1.11), pyruvate kinase (EC 2.7.1.40), enzymes from purine salvage pathway, purine catabolism, and stringent response factor Rel (EC 2.7.6.5).
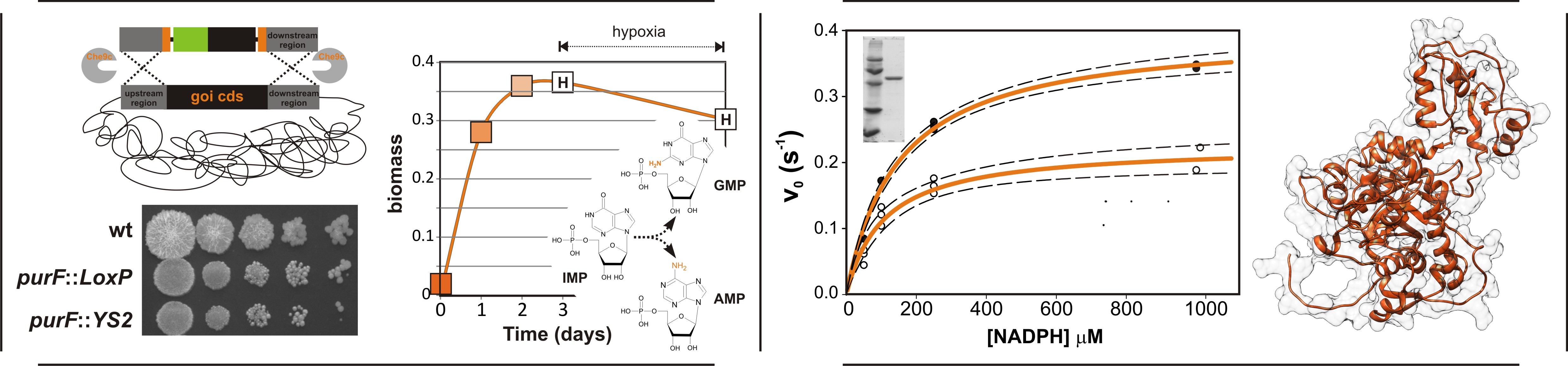
We investigate the impacts of the variations of different conditions on the function of studied enzymes. Among those are conditions mimicking an everchanging intracellular environment, fluctuation in the levels of metabolites and interactions with cellular proteins. We use nonpathogenic Mycobacterium smegmatis as a model organism for our bacterial studies. By generating knock-out mutants of enzymes from both the de novo and salvage pathways of purine metabolism, we aim to identify the key enzymes influencing the biosynthesis of different bases and use them as targets to develop specific inhibitors. Other approaches include protein-protein interaction studies, cultivation assays or proteomics/metabolomics to understand the complex regulation of the mycobacterial metabolism.
Our publications (2014-)
- Knejzlík Z., Doležal M., Herkommerová K., Clarova K., Klíma M., Dedola M., Zborníková E., Rejman D., Pichová I.: The mycobacterial guaB1 gene encodes a guanosine 5′-monophosphate reductase with a cystathionine-β-synthase domain. 2022, 289:5571-5598.
- Snášel J., Machová I., Šolínová V., Kašička V., Krečmerová M., Pichová I.: Phosphofructokinases A and B from Mycobacterium tuberculosis Display Different Catalytic Properties and Allosteric Regulation. J. Int. J. Mol. Sci. 2021, 22(3):1483.
- Knejzlík Z., Herkommerová K., Hocková D., Pichová I.: Hypoxanthine-Guanine Phosphoribosyltransferase Is Dispensable for Mycobacterium smegmatis Viability. J. Bacteriol. 2020, 202(5):e00710-19.
- Knejzlík Z., Herkommerová K., Pichová I.: Catabolism of 8-oxo-purines is mainly routed via the guanine to xanthine interconversion pathway in Mycobacterium smegmatis. Tuberculosis. 2019, 119:101879.
- Zborníková E., Knejzlík Z., Hauryliuk V., Krásný L., Rejman D.: Analysis of nucleotide pools in bacteria using HPLC-MS in HILIC mode. Talanta. 2019, 205:120161.
- Snášel J., Pichová I.: Allosteric regulation of pyruvate kinase from Mycobacterium tuberculosis by metabolites. Biochim. Biophys. Acta-Prot. Proteom. 2019, 1867(2):125-139.
- Machová I., Hubálek M., Lepšík M., Bednárová L., Pazderková M., Kopecký V. Jr., Snášel J., Dostál J., Pichová I.: The Role of Cysteine Residues in Catalysis of Phosphoenolpyruvate Carboxykinase from Mycobacterium tuberculosis. PLoS One. 2017, 12(1):e0170373.
- Machová I., Snášel J., Dostál J., Brynda J., Fanfrlík J., Singh M., Tarábek J., Vaněk O., Bednárová L., Pichová I.: Structural and functional studies of phosphoenolpyruvate carboxykinase from Mycobacterium tuberculosis. PLoS One. 2015, 10(3):e0120682.
- Snášel J., Nauš P., Dostál J., Hnízda A., Fanfrlík J., Brynda J., Bourderioux A., Dušek M., Dvořáková H., Stolaříková J., Zábranská H., Pohl R., Konečný P., Džubák P., Votruba I., Hajdúch M., Řezáčová P., Veverka V., Hocek M., Pichová I.: Structural Basis for Inhibition of Mycobacterial and Human Adenosine Kinase by 7-Substituted 7-(Het)aryl-7-deazaadenine Ribonucleosides. J. Med. Chem. 2014, 57(20):8268-8279.
- Machová I., Snašel J., Zimmermann M., Laubitz D., Plocinski P., Oehlmann W., Singh M., Dostál J., Sauer U., Pichová I.: Mycobacterium tuberculosis phosphoenolpyruvate carboxykinase is regulated by redox mechanisms and interaction with thioredoxin. J. Biol. Chem. 2014, 289(19):13066-78.
People
Zdeněk Knejzlík, Jan Snášel, Michal Doležal, Ondřej Bulvas, Matteo Dedola, Kamila Clarová, Dagmar Grundová, Elena Dolejší

